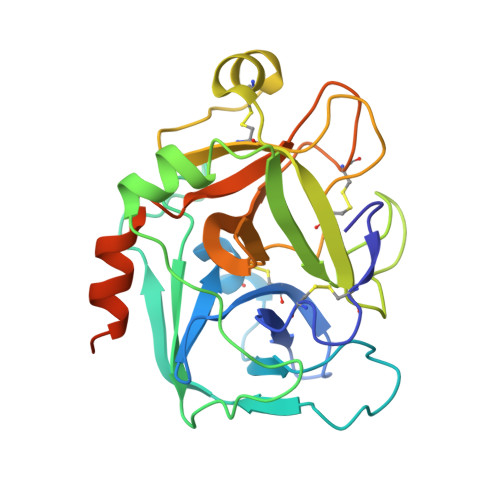
Arylsulfonamides: A Study of the Relationship between Activity and Conformational Preferences for a Series of Factor Xa Inhibitors.
Senger, S., Convery, M.A., Chan, C., Watson, N.S.(2006) Bioorg Med Chem Lett 16: 5731
- PubMed: 16982192
- DOI: https://doi.org/10.1016/j.bmcl.2006.08.092
- Primary Citation of Related Structures:
2J2U, 2J34, 2J38 - PubMed Abstract:
Torsional scans of sulfonamide S-C bonds in small model systems of a series of arylsulfonamide factor Xa inhibitors were performed in order to investigate if conformational effects can help to rationalise the observed SAR. Computational results were in good agreement with the experimental data indicating that the sulfonamide conformation plays an important role in determining the activity in this particular series of factor Xa inhibitors.
Organizational Affiliation:
GlaxoSmithKline, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire SG1 2NY, UK. stefan.x.senger@gsk.com